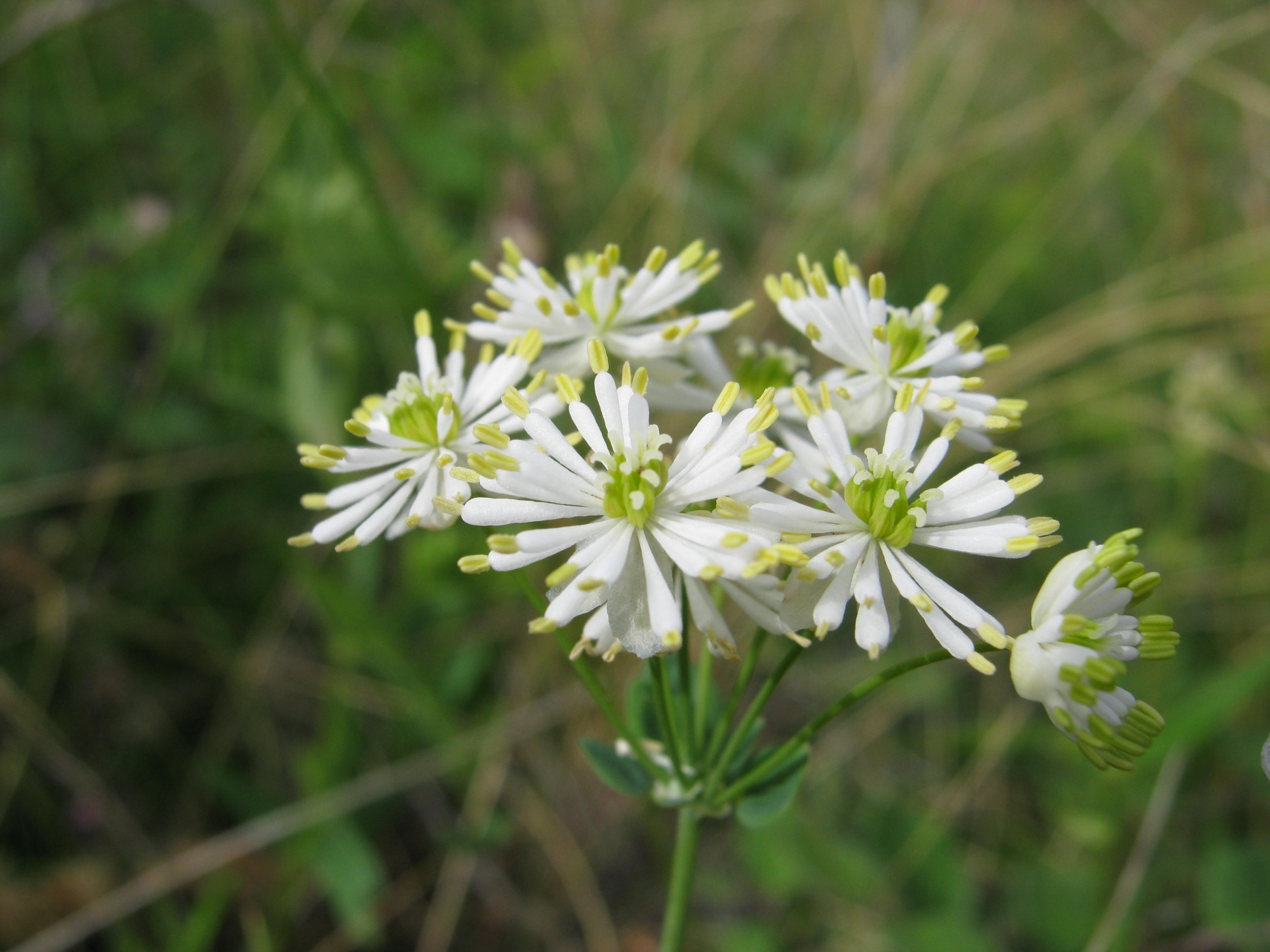
“Next generation sequencing provides us with a large amount of data on plant genomes, proteomes, transcriptomes, etc. Numerical techniques to make sense of these data are provided by a newly emerging field of botany – plant bioinformatics. We used bioinformatics technologies to study the Thalictrum genus,” says Andrey Erst, senior researcher at the TSU laboratory Herbarium and co-author of the article.
The Russian-Chinese research team has also analyzed selective pressure and codon usage bias of all the plastid coding genes for 11 species. Phylogenetic relationships showed that Thalictrum is a monophyly and divided into two major clades. The availability of data on these plastomes offers valuable genetic information for accurate identification of species and taxonomy, phylogenetic resolution, and evolutionary studies of Thalictrum, and should assist in exploring and using Thalictrum plants.“It’s a phylogenetically and economically important member of the family, but it is also considered one of the hardest plants in which to settle phylogenetic and taxonomic questions between taxons. We were able to identify plastome features that were mostly conserved in the overall structure of the gene order: a quadripartite structure, loci locations, and gene structure. We have also discovered eight highly variable noncoding genome regions. They can be used for a phylogeographic and taxon analysis of various species,” says Andrey Erst.